R - Data visualization, linear regression and logistic regression
Welcome. In this post, I am going to show you how you may create point maps, linear regression and logistic regression utilizing R base tools and tidymodels workflow.
 Image credit: Pott L.P.
Image credit: Pott L.P.Data visualization, linear regression and logistic regression
Crop data collection - ground truth and remote sensing
Loading the packages
library(tidyverse)
library(tidymodels)
library(sf)
library(geobr)URL link of data from data collection and remote sensing on GitHub https://github.com/luanpott10/Class
You will need the raw file https://raw.githubusercontent.com/luanpott10/Class/main/data_crops.csv
Loading the data
data <- read.csv("https://raw.githubusercontent.com/luanpott10/Class/main/data_crops.csv")Plot the data in the map
cities_RS <- read_municipality(code_muni = "RS", year= 2020)
ggplot()+
geom_sf(data=cities_RS)+
geom_point(data=data,aes(x=longitude,y=latitude,fill=class),shape=22,size=2)+
labs(x= "Longitude", y = "Latitude")+
scale_fill_manual(values = c("#eded0c", "#49a345"))+
theme(legend.position = c(0.17, 0.2),
panel.border = element_rect(color="Black", fill = NA),
panel.background = element_rect(fill = "#f2f2f2"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.title.x = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.x = element_text(family = "serif",
colour = "#000000",size = 18.0),
axis.title.y = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.y = element_text(family = "serif",
colour = "#000000",size = 18.0),
legend.title = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.text = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.background = element_rect(fill="#f2f2f2",
linetype="dashed",
colour ="#f2f2f2"))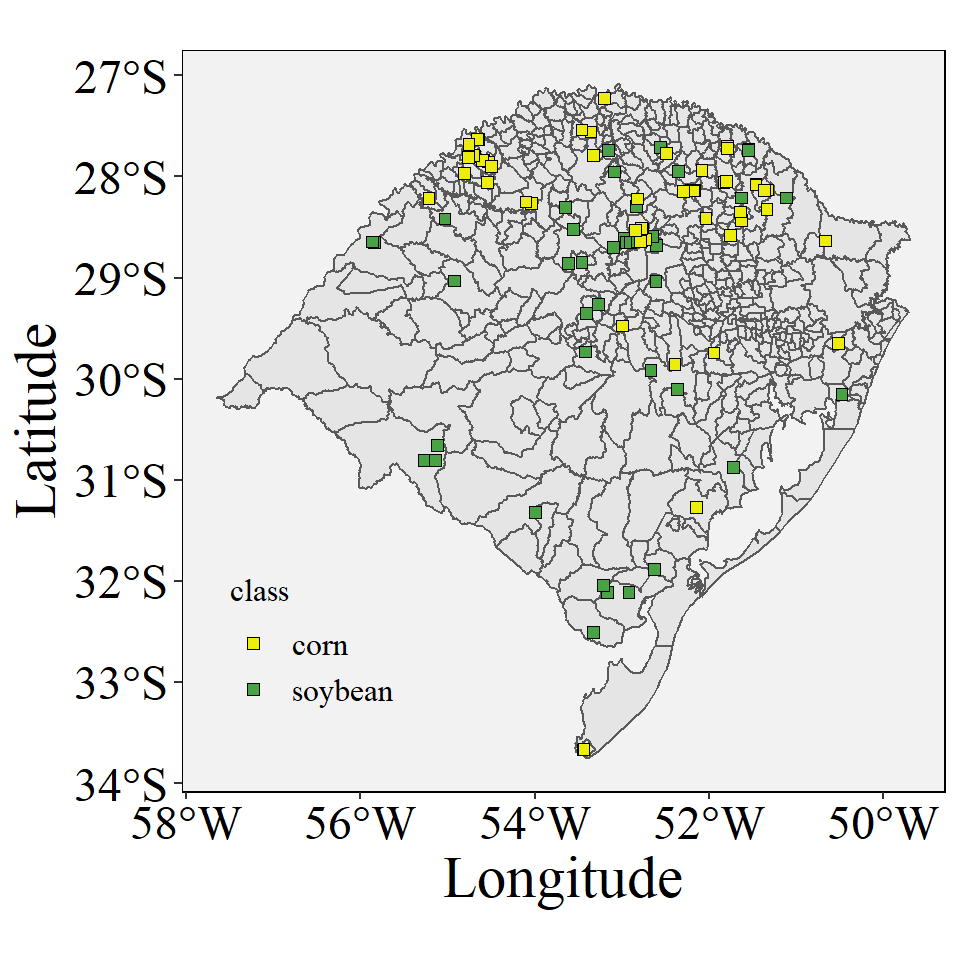
Plot the data variables - continuous x continuous
ggplot(data=data,aes(x=b3_GCVI,y=b4_GCVI))+
geom_point(aes(fill=class),shape=22,size=2)+
scale_fill_manual(values = c("#eded0c", "#49a345"))+
stat_smooth(formula = y~x, method="lm", se=FALSE,color="black", linetype='dashed')+
theme(panel.border = element_rect(color="Black", fill = NA),
panel.background = element_rect(fill = "#f2f2f2"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.title.x = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.x = element_text(family = "serif",
colour = "#000000",size = 18.0),
axis.title.y = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.y = element_text(family = "serif",
colour = "#000000",size = 18.0),
legend.title = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.text = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.background = element_rect(fill="#f2f2f2",
linetype="dashed",
colour ="#f2f2f2"))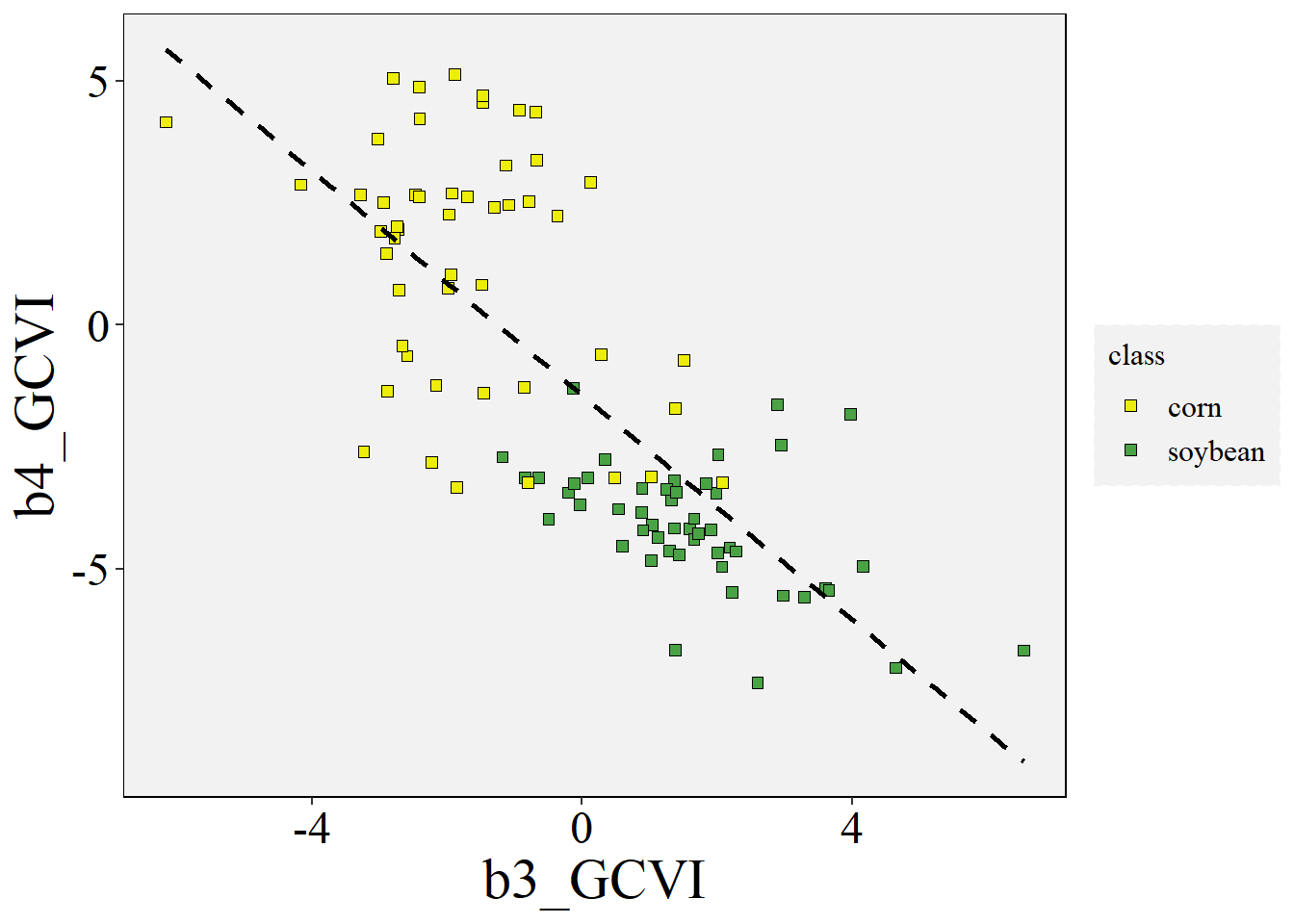
Linear regression
lm_fit_x <- lm(b3_GCVI ~ b4_GCVI, data = data)
summary(lm_fit_x)
##
## Call:
## lm(formula = b3_GCVI ~ b4_GCVI, data = data)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.7924 -0.9983 -0.0018 0.9525 3.9586
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -0.73701 0.15933 -4.626 1.14e-05 ***
## b4_GCVI -0.49793 0.04352 -11.440 < 2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.475 on 98 degrees of freedom
## Multiple R-squared: 0.5718, Adjusted R-squared: 0.5675
## F-statistic: 130.9 on 1 and 98 DF, p-value: < 2.2e-16Linear regression by tidymodels workflow
Creating a parsnip specification for a linear regression model
lm_model <- linear_reg() |>
set_engine('lm') |>
set_mode('regression')Fitting the model supplying a formula expression and the data
lm_fit <- lm_model %>%
fit(b3_GCVI ~ b4_GCVI, data = data)Summary of the model
lm_fit |>
pluck("fit") |>
summary()
##
## Call:
## stats::lm(formula = b3_GCVI ~ b4_GCVI, data = data)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.7924 -0.9983 -0.0018 0.9525 3.9586
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -0.73701 0.15933 -4.626 1.14e-05 ***
## b4_GCVI -0.49793 0.04352 -11.440 < 2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.475 on 98 degrees of freedom
## Multiple R-squared: 0.5718, Adjusted R-squared: 0.5675
## F-statistic: 130.9 on 1 and 98 DF, p-value: < 2.2e-16
# Also you can use
lm_fit$fit |> summary()
##
## Call:
## stats::lm(formula = b3_GCVI ~ b4_GCVI, data = data)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.7924 -0.9983 -0.0018 0.9525 3.9586
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -0.73701 0.15933 -4.626 1.14e-05 ***
## b4_GCVI -0.49793 0.04352 -11.440 < 2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.475 on 98 degrees of freedom
## Multiple R-squared: 0.5718, Adjusted R-squared: 0.5675
## F-statistic: 130.9 on 1 and 98 DF, p-value: < 2.2e-16Parameter estimates of a the lm object
tidy(lm_fit)
## # A tibble: 2 x 5
## term estimate std.error statistic p.value
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) -0.737 0.159 -4.63 1.14e- 5
## 2 b4_GCVI -0.498 0.0435 -11.4 9.39e-20Extract the model statistics
glance(lm_fit)
## # A tibble: 1 x 12
## r.squared adj.r.squared sigma statistic p.value df logLik AIC BIC
## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 0.572 0.567 1.47 131. 9.39e-20 1 -180. 365. 373.
## # ... with 3 more variables: deviance <dbl>, df.residual <int>, nobs <int>Plot the data variables - categorical x continuous
data <- data |> mutate(target = case_when(class == "corn" ~ 0,
class == "soybean" ~ 1))
ggplot(data=data, aes(y=target,x=b4_GCVI))+
geom_point(aes(fill=as.factor(target)),shape=22,size=2) +
scale_fill_manual(values = c("#eded0c", "#49a345"))+
scale_y_continuous(breaks=c(0,1),
labels=c("0","1"),
limits=c(0,1))+
stat_smooth(formula = y~x, method="glm", se=FALSE, method.args = list(family=binomial),
color="black", linetype='dashed')+
labs(x= "b4_GCVI", y = "Class",fill="Class")+
theme(legend.position = c(0.17, 0.2),
panel.border = element_rect(color="Black", fill = NA),
panel.background = element_rect(fill = "#f2f2f2"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.title.x = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.x = element_text(family = "serif",
colour = "#000000",size = 18.0),
axis.title.y = element_text(family = "serif",
colour = "#000000",size = 22.0),
axis.text.y = element_text(family = "serif",
colour = "#000000",size = 18.0),
legend.title = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.text = element_text(family = "serif",
colour = '#000000',size = 12.0),
legend.background = element_rect(fill="#f2f2f2",
linetype="dashed",
colour ="#f2f2f2"))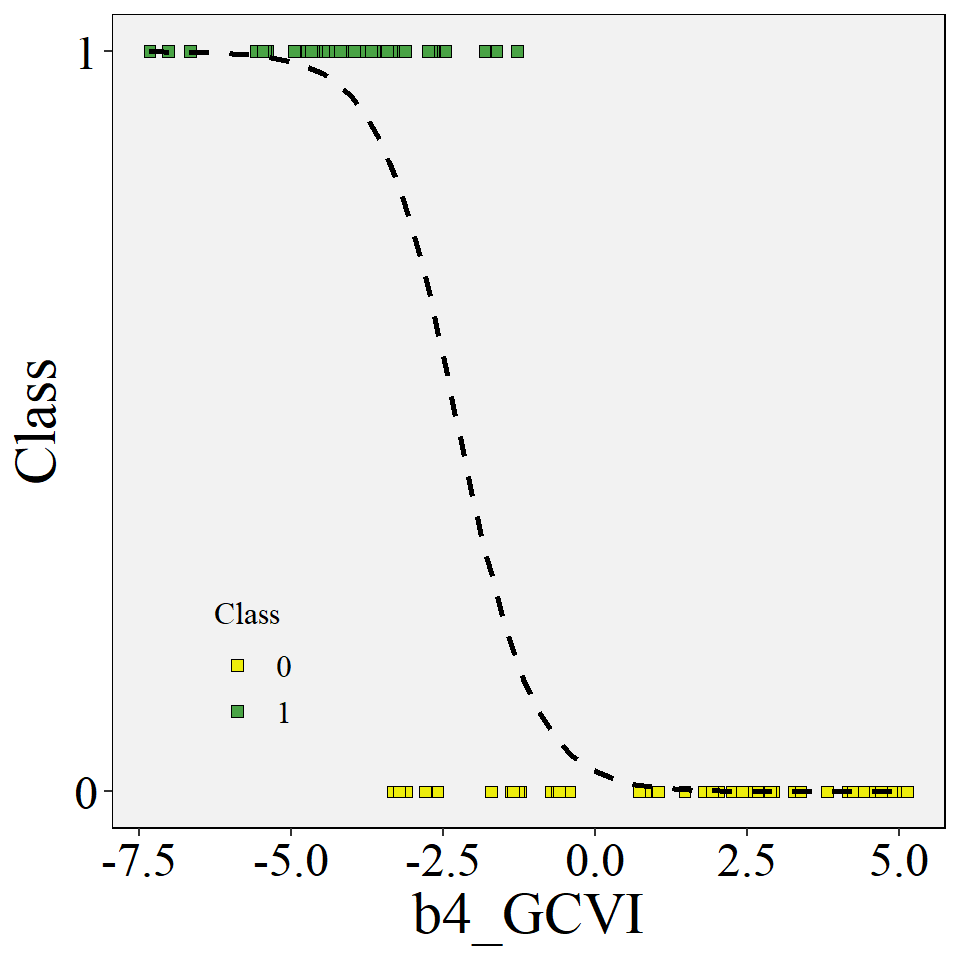
Logistic regression
lg_fit_x <- glm(as.factor(class) ~ b4_GCVI, family="binomial", data=data)
summary(lg_fit_x)
##
## Call:
## glm(formula = as.factor(class) ~ b4_GCVI, family = "binomial",
## data = data)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.91207 -0.04298 0.01134 0.32227 1.86651
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -3.5734 1.1155 -3.203 0.00136 **
## b4_GCVI -1.5677 0.3778 -4.150 3.33e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 138.629 on 99 degrees of freedom
## Residual deviance: 42.379 on 98 degrees of freedom
## AIC: 46.379
##
## Number of Fisher Scoring iterations: 7Logistic regression by tidymodels workflow
Creating a parsnip specification for a logistic regression model
lg_model <- logistic_reg() |>
set_engine("glm") |>
set_mode("classification")Fitting the model supplying a formula expression and the data
lg_fit <- lg_model |>
fit(as.factor(class) ~ b4_GCVI, data = data)Summary of the model
lg_fit |>
pluck("fit") |>
summary()
##
## Call:
## stats::glm(formula = as.factor(class) ~ b4_GCVI, family = stats::binomial,
## data = data)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.91207 -0.04298 0.01134 0.32227 1.86651
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -3.5734 1.1155 -3.203 0.00136 **
## b4_GCVI -1.5677 0.3778 -4.150 3.33e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 138.629 on 99 degrees of freedom
## Residual deviance: 42.379 on 98 degrees of freedom
## AIC: 46.379
##
## Number of Fisher Scoring iterations: 7
# Also you can use
lg_fit$fit |> summary()
##
## Call:
## stats::glm(formula = as.factor(class) ~ b4_GCVI, family = stats::binomial,
## data = data)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.91207 -0.04298 0.01134 0.32227 1.86651
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -3.5734 1.1155 -3.203 0.00136 **
## b4_GCVI -1.5677 0.3778 -4.150 3.33e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 138.629 on 99 degrees of freedom
## Residual deviance: 42.379 on 98 degrees of freedom
## AIC: 46.379
##
## Number of Fisher Scoring iterations: 7Parameter estimates of a the lm object
tidy(lg_fit)
## # A tibble: 2 x 5
## term estimate std.error statistic p.value
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) -3.57 1.12 -3.20 0.00136
## 2 b4_GCVI -1.57 0.378 -4.15 0.0000333Extract the model statistics
glance(lg_fit)
## # A tibble: 1 x 8
## null.deviance df.null logLik AIC BIC deviance df.residual nobs
## <dbl> <int> <dbl> <dbl> <dbl> <dbl> <int> <int>
## 1 139. 99 -21.2 46.4 51.6 42.4 98 100